When it comes to the architecture of the human genome, it’s only a matter of time before harmful genes—genes that could compromise future generations—arise in a population. These mutations accumulate in the gene pool, primarily affected by a population’s size and practices like marrying within a small community, according to researchers.
But much of the information about the effects of a population’s mutation load is based on genetic theory, with limited direct evidence concerning the effects on evolutionary fitness, or fertility.
New research from University of California, Davis, provides rare direct evidence showing that increased homozygosity—meaning two identical alleles in a genome—leads to negative effects on fertility in a human population. The paper was published Oct. 17 in the Proceedings of the National Academy of Sciences journal.
“People have known since Darwin that if you take people who are first cousins and they have children together, the children are more likely to develop certain diseases or be less healthy,” said Brenna Henn, an associate professor of anthropology in the College of Letters and Science at UC Davis.
The research assesses the consequences of homozygosity among Namibia’s Himba community, an isolated, agro-pastoralist population in which marriage between people with the same ancestor occurs. The research was led by Natalie Swinford, who received her doctoral degree in 2022 in evolutionary anthropology and human population genetics, and Henn.
“They’re what we call an ‘endogamous population,’ meaning people are meeting their partners just from within that Himba group,” said Henn. “They also have a unique system of marriage and reproduction, where men and women can have multiple boyfriends or girlfriends during their marriage. That means there are a lot of half-siblings in the population. That’s a unique feature and it means that we can leverage that social structure to look at different genetic effects.”
Echoes in the genome
In the study, the team gathered genetic data from 681 individuals from the Himba population. Genetic analyses revealed that the Himba have genetic markers that show higher levels of inbreeding.
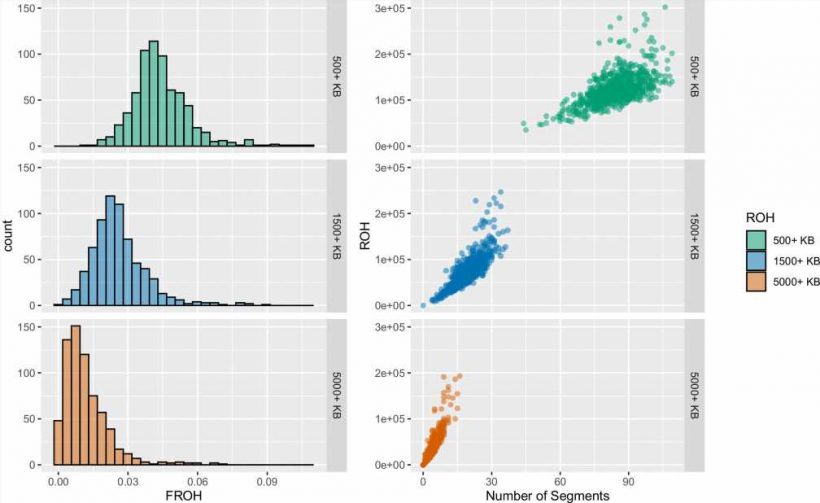
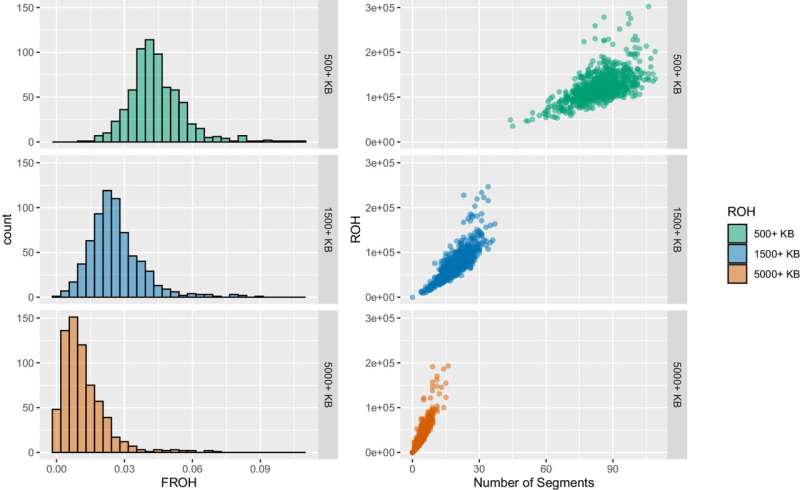
Known as runs of homozygosity, or ROH, these markers are multiple and particularly long in the genomes of the analyzed Himba, which indicates that their parents had a high likelihood of sharing an ancestor.
While the Himba population has historically exhibited a preference for consanguinity, Henn and Swinford were surprised to find that none of the individuals in their sample population had parents who were actually first cousins. The lengths of the ROH in the genomes indicated otherwise.
The researchers found that these genetic effects can pool and accumulate over time. So, bottleneck events, like a decrease in population that leads to inbreeding, can have genetic echoes that don’t manifest until generations later. The researchers concluded that such events occurred within the past 12 to 18 generations of the Himba population.
“People may not be full first-cousins,” Henn said. “But they may be half-cousins once removed and then their grandparents might have been half-cousins. Anytime something like that happens, it’s going to contribute to there being identical DNA in the offspring.”
A negative effect on fertility
The Himba are a pronatalist community, meaning they encourage their members to have many children. Typically, there are short intervals between births, roughly between one and three years, researchers said.
To gauge the effects of long ROH on fertility, the researchers measured the reproductive success of post-reproductive women in their sample population. The researchers defined reproductive success as the number of children who survived to at least 5.
The research team used statistical models to analyze the relationship between the amount of ROH in the genome and the number of children a woman had. They found that the greater the proportion of the genome that was in ROH, the more likely a woman was to have fewer children than a woman who had less ROH.
“This means that a woman who has parents who are more related is more likely to have fewer children throughout her lifetime than a woman who has parents who are less related,” said Swinford.
Contributing authors on the paper include S.P. Prall, University of Missouri; S. Gopalan, Duke University; C.M. Williams, Brown University; J. Sheehama, University of Namibia; and B.A. Scelza, University of California, Los Angeles.
More information:
N. A. Swinford et al, Increased homozygosity due to endogamy results in fitness consequences in a human population, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2309552120
Journal information:
Proceedings of the National Academy of Sciences
Source: Read Full Article