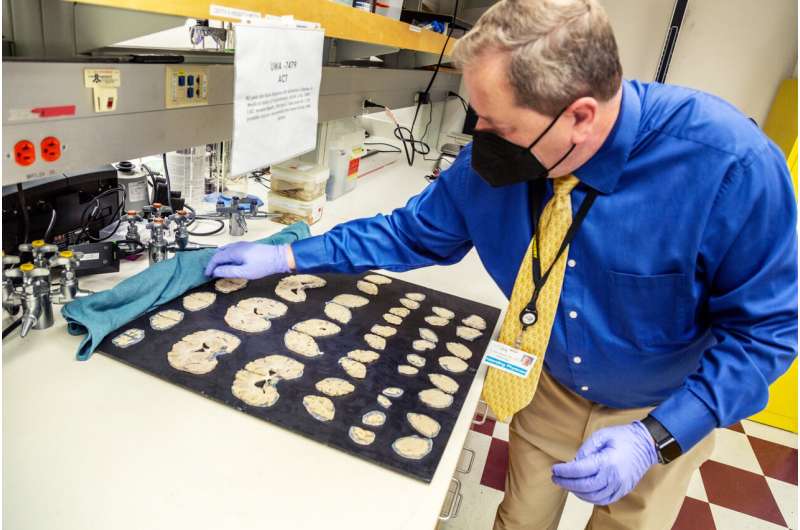
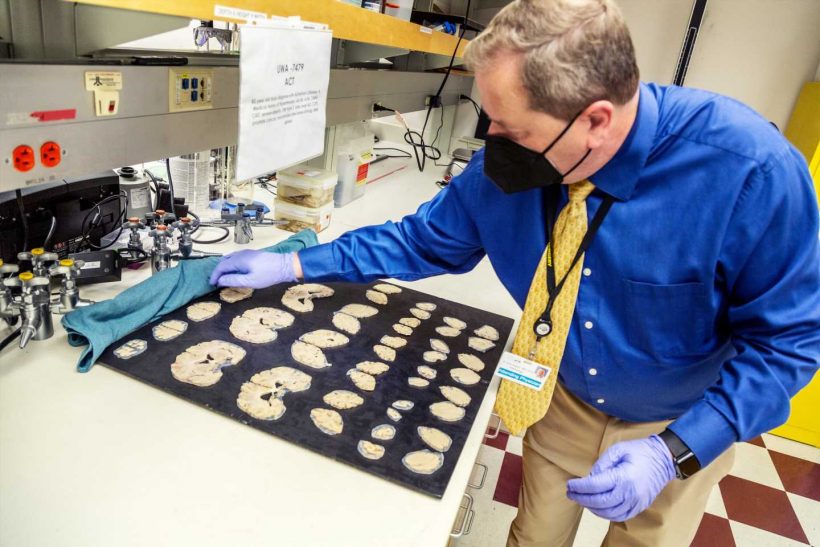
If you compare the brain of someone who has died from neurodegenerative disease to that of a healthy person, you can’t miss the difference: In the case of severe Alzheimer’s, the brain will be noticeably smaller, with large gaps where pieces would normally nestle close together.
This shrinkage, termed brain atrophy, is due to the die-off of neurons and their connections across the entire cortex, the wrinkled, outermost shell of the brain. Nobody knows what triggers these huge brain changes in Alzheimer’s disease, let alone how to stop or reverse them.
But neuroscientists at the Allen Institute and their collaborators are focusing on changes at a much smaller, granular scale, hoping to provide incredibly detailed insight into what exactly goes wrong in Alzheimer’s disease. They’ve just publicly released some of the first data showing the specific types of neurons and other brain cells that die off or otherwise change in Alzheimer’s disease, using cutting-edge techniques to categorize individual cells based on what their genes do. This approach could ultimately identify new targets for better therapies to slow or halt the disease’s progression by preventing these specific cell populations from dying.
This method of mapping a diseased brain in cellular detail is only recently possible, thanks to new techniques to study large numbers of individual brain cells, said Ed Lein, Ph.D., Senior Investigator at the Allen Institute for Brain Science, a division of the Allen Institute, and lead investigator of the Seattle Alzheimer’s Disease Brain Cell Atlas team, which released the new data.
“We’ve made really remarkable progress in developing new tools to study the human brain, and this really opens up whole new possibilities for studying disease,” Lein said. “As we’ve developed these tools, it became apparent that we could have a big impact by creating a much higher resolution atlas of what Alzheimer’s actually looks like at the cellular level.”
The basics of Alzheimer
The Seattle Alzheimer’s Disease Brain Cell Atlas consortium, or SEA-AD, is a National Institute on Aging-funded collaboration headquartered at the Allen Institute, with additional research projects at UW Medicine and Kaiser Permanente Washington Health Research Institute. The publicly available dataset captures large-scale cellular and molecular information gleaned from more than 1.2 million neurons and other brain cells from 84 people who donated their brains to science after their deaths, as well as detailed microscopy images of amyloid-β and other disease-related proteins in their brains.
The cellular techniques used in the consortium build off previous work at the Allen Institute and elsewhere, through the NIH-funded BRAIN Initiative, that uses genes switched on in individual brain cells to categorize them into discrete types. These methods were first used to understand the basic cellular building blocks of a healthy brain, and they are now being turned to understand Alzheimer’s disease at a new level of detail and resolution.
“Because we don’t know yet which of many possible pathways are most important in Alzheimer’s, we need to have the most comprehensive picture we possibly can of the related changes in brain cells,” said Richard J. Hodes, M.D., Director of NIH’s National Institute on Aging. “Progress such as this means we have an increased understanding of what underlies the disease, and therefore a better hope for strategizing to effectively prevent and treat it.”
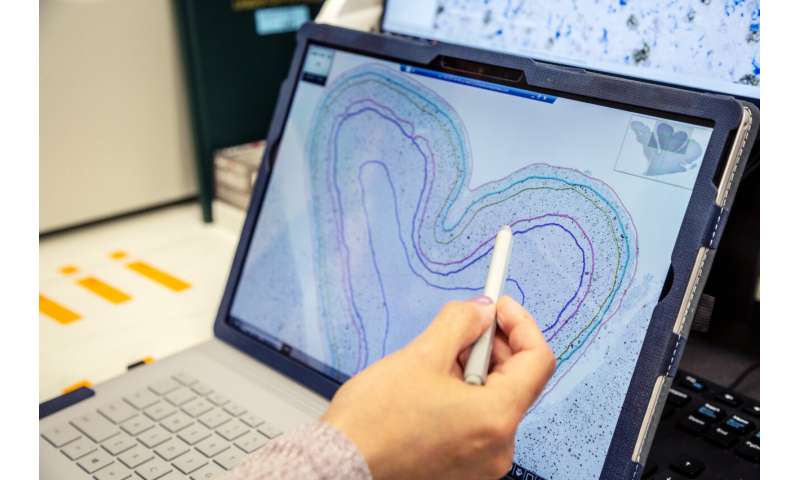

Scientists still don’t understand the biological changes that trigger Alzheimer’s disease, which afflicts more than 6 million Americans and is the most common cause of dementia. For decades, scientists thought amyloid plaques—large clumps of a naturally occurring protein, amyloid-β, which is toxic to neurons at high levels—was the root cause of Alzheimer’s. But several recently developed drugs that break up amyloid plaques have had minimal, if any, effect on disease progression. The SEA-AD team wanted to go back to the basics, identifying the individual kinds of neurons and other brain cells that die or change as Alzheimer’s progresses, with the ultimate goal of developing targeted therapies to protect these cells before they die.
In this first data release, which analyzes cells from one region of the brain, the team identified certain kinds of neurons that are selectively vulnerable to the disease, while other kinds of brain cells increase in abundance. Understanding the true cellular roots of the disease will take more data, from more parts of the brain.
“This data release is the first of many. My hope is that with this release and future releases, we will generate data that will give us clues as to how this disease actually works,” said C. Dirk Keene, M.D., Ph.D., Professor and Nancy and Buster Alvord Endowed Chair of Neuropathology at UW Medicine and one of the investigators involved in the data release. “The more scientists that take innovative approaches to try to understand the disease, the more likely it is that we’re going to find those drug targets and develop drugs that will be effective.”
The Alzheimer’s brain parts list
Alzheimer’s disease doesn’t affect the entire brain at once. The disease begins in a region of the brain involved in memory called the entorhinal cortex and starts to affect nearby parts of the brain as the disease progresses, eventually killing neurons across the entire cortex. Eventually, the Seattle researchers want to understand the entire progression of the disease, but for their first data release, they focused on a region of the cortex known as the middle temporal gyrus, which is affected approximately midway in the course of Alzheimer’s progression.
The 84 people who donated their brains for this research represent a spectrum across all stages of Alzheimer’s disease, from healthy aging to severe dementia, and are all participants in the Adult Changes in Thought (ACT) study, a long-running study of brain aging led by Kaiser Permanente Washington and UW Medicine, or studies in the UW Medicine Alzheimer’s Disease Research Center. Brain donations are coordinated through the University of Washington School of Medicine BioRepository and Integrated Research (BRaIN) laboratory, directed by Keene, who describes brain donation as “the greatest gift one can give to science, without which this research would not be possible.”
The scientists used a variety of molecular characteristics, including assays to identify the genes switched on in individual cells in a method known as single-cell RNA sequencing, to categorize more than 1.2 million brain cells from the 84 donors into individual cell types. The scientists compared these cell types to those from younger, healthy brain donors—a cell-type reference map generated earlier by the Allen Institute team and their collaborators through the NIH-funded BRAIN Initiative Cell Census Network. Together with the single-cell assays, the team also conducts highly quantitative neuropathology of the brain region to better understand the spatial makeup of disease-related proteins and specific types of cells in the donors’ brains.
Delving into the data, the scientists found some intriguing differences in the Alzheimer’s patients’ cells. For example, they found that some of the neurons selectively vulnerable to Alzheimer’s are those that make long-range connections (think highways instead of streets) across the cortex, the brain’s center for more complex cognitive functions. The loss of these particular neurons aligns with the disease’s effects: Alzheimer’s patients primarily lose cognitive abilities like memory, language and learning. They also saw increases in the proportions of certain non-neuronal brain cells, including microglia, which act as immune cells of the brain.
“Now that we see these specific neuron populations falling out, the big question is why?” said Kyle Travaglini, Ph.D., a scientist at the Allen Institute in the SEA-AD group who will present findings from the data release on Tuesday, Aug. 2, at the Alzheimer’s Association International Conference in San Diego. “The vision is to identify a vulnerable population and create a therapy to protect it and prevent it from dying. In order to get there, we need to understand why these neurons are dying in the first place.”
The scientists hope that sifting through differences in how genes are switched on or off in Alzheimer’s disease will start to get at that question. They’re also planning to survey many more regions of the brain to understand the full course Alzheimer’s takes in its destructive progression.
“We’re not going to solve this by looking at one brain region,” Lein said. “We want to describe trajectories of the disease as they march across brain regions and across different cell types in those brain regions, including regions affected both early and late in the disease. The aim, ultimately, is to find the early, causal events that happen when the disease is still potentially reversible.”
The data release opens SEA-AD data and resources to the general scientific community for anyone to explore and includes:
- A transcriptomics comparative viewer, which captures side-by-side changes in gene activity in single cells between the healthy, reference brains and brains from the cohort of 84 donors across the spectrum of Alzheimer’s disease. The viewer can be filtered by cell type or by several different donor characteristics, including age at death, sex, presence of dementia, and years of education.
- A transcriptomic explorer that shows the reference set of MTG brain cell types from the healthy, reference donors. This data visualization shows the relationships between all the cell types in this region, as well as expression levels for any genes of interest.
- A donor index and neuropathology image viewer, which captures demographic, clinical, cognitive and pathological information from the 84 brain donors and detailed pathology images and quantifications from each brain region. The UW Medicine pathology team developed machine learning methods to precisely quantify levels of disease-related proteins in these images.
- A link to the SEA-AD transcriptomics data in the Chan Zuckerberg CELL by GENE, an interactive data explorer for single-cell datasets.
- A link to the SEA-AD chromatin accessibility data in the UC Santa Cruz’s Genome Browser.
Source: Read Full Article
