Several SARS-CoV-2 variants of concern have emerged since the start of the COVID-19 pandemic. Each bear mutations that may enhance transmissibility, allow SARS-CoV-2 to escape established immunity or otherwise alter the virus sufficiently to generate novel characteristics.
The SARS-CoV-2 delta variant bears the D164G mutation, which is known to enhance viral fitness. However, a number of additional mutations are known to be common to the delta and delta plus lineages, at least 25 as identified in a study recently published in the Journal of Autoimmunity, mainly located on the spike protein of the virus.
.jpg)
The delta and delta plus variants are distinct
Within this report, mutations T95I and W258L were found to be present in 40% of delta plus genomes, with the latter lacking from the delta genome. The authors propose that these should be included in defining the characteristic mutations of the delta and delta plus variants.
In search of other defining mutations, the group analyzed high-quality genome data collated in the global science initiative and primary source database, finding 656 unique mutations to the delta variant compared with wildtype and 269 to the delta plus variant.
High prevalence mutations were present in at least 20% of samples. However, they were greater amongst the delta plus variant, with 40 such mutations compared with 29. Two mutations, in particular, were heavily prevalent amongst the delta plus variant and almost absent from the delta variant: V70F and W258L.
Mutations A222V and T95I to the spike protein were present amongst 58% and 38% of delta plus variants, respectively, with the same mutations present in only 9% and 22% of delta variants. Mutations to regions other than the spike protein followed a similar trend, with mutation A328T to non-structural protein 3 being present in 58% of delta plus samples and none of the delta variants. Other mutations to non-structural proteins such as A446V (nsp4) are more strongly present amongst the delta plus variant, 58% compared to 9%.
The delta plus variant has largely been distinguished from the delta variant by the presence of the K417N mutation, though this analysis implies that several additional mutations must be considered.
The group next performed relative abundance analysis to correlate the occurrence of mutations with more than 20% abundance in each strain. Within the delta variant, all mutations except T95I and G142D co-occurred in all cases, with these mutations being present in only ~25% and ~50% of cases bearing other mutations, respectively.
Within the delta plus variant, 40% of sequences contained the W258L mutation, amongst which there was a strong correlation with all other mutations, including the aforementioned G142D and T95I mutations to the spike protein. Amongst the delta plus variant samples bearing the signature delta variant mutation D950N, mutation A446V to nsp4 was present in 90% of cases.
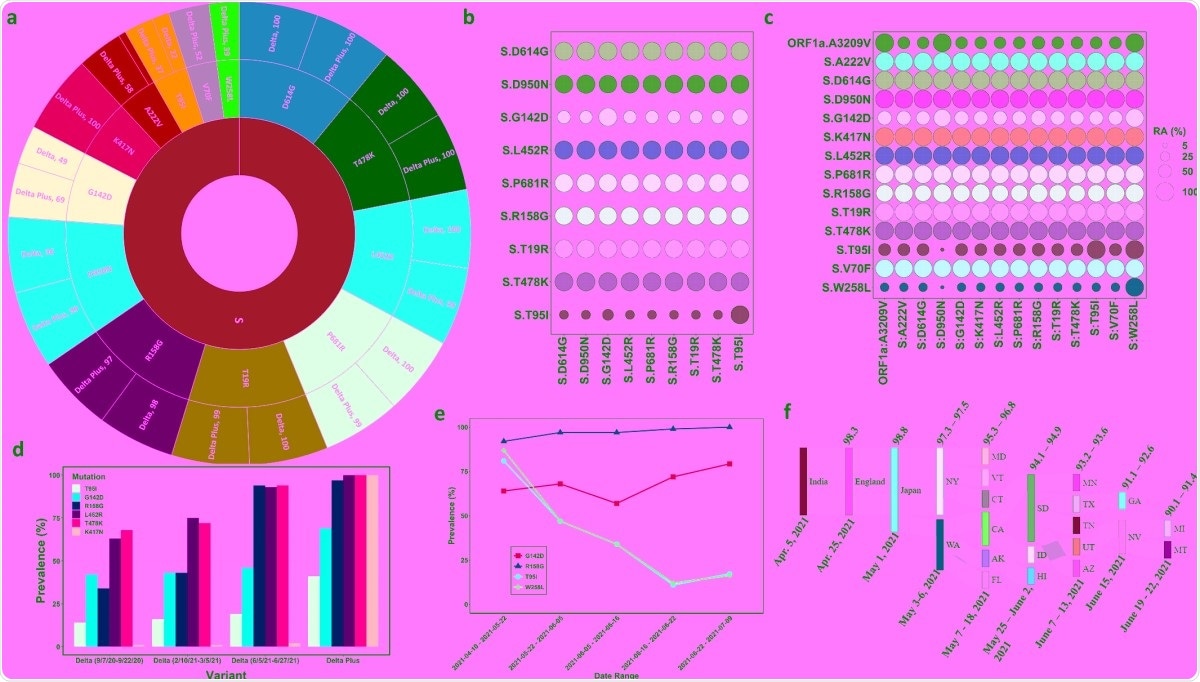
The acquisition of mutations by the delta lineage
To study the dynamics of the key mutations (T95I, G142D, R158G, L452R, T478K, and K417N) in each strain, the group determined their prevalence at several time points. They found that each increased in presence within the delta strain over the course of a year, and that each was notably more present in the delta plus strain since its origin.
The K478 mutation of the delta plus variant is known to enhance immune evasion towards some antibodies thanks to an extended side chain that impedes binding, resulting in greater transmissibility of the strain. The influence of other delta plus mutations is less well explained, and thus the group examined cryo-electron microscopy images to determine what structural changes the mutations may induce.
The G142D mutation, which frequently co-occurred with other mutations, was found to cause a steric clash with sidechain R158 and disrupt the conformation of the spike protein. Mutation R158G eliminates this steric hindrance, and this mutation was found in combination with G142D in all delta plus cases. Another mutation, K417N, was noted to act similarly to mutations to K478, hindering the binding of antibodies to the spike protein.
When tracking the path of the delta variant across the globe, the group state that the strain likely originated in India and traveled through the UK, Japan, and eventually Washington, USA. Since then, the strain has adopted key mutations to generate the distinct delta plus variant.
The group highlights the large number of distinct mutations characteristic of this strain that frequently co-occur. At this point, the delta and delta plus variants have diversified with unique mutation profiles, somewhat fuelled by pockets of local infection, giving each a chance to diverge.
- Kannan et al. (2021) Evolutionary analysis of the Delta and Delta Plus variants of the SARS-CoV-2 viruses. Journal of Autoimmunity. https://doi.org/10.1016/j.jaut.2021.102715, https://www.sciencedirect.com/science/article/pii/S0896841121001232?via%3Dihub.
Posted in: Medical Science News | Medical Research News | Disease/Infection News
Tags: Antibodies, Autoimmunity, Coronavirus Disease COVID-19, Electron, Electron Microscopy, Evolution, Genome, immunity, Microscopy, Mutation, Pandemic, Protein, SARS, SARS-CoV-2, Spike Protein, Structural Protein, Virus

Written by
Michael Greenwood
Michael graduated from Manchester Metropolitan University with a B.Sc. in Chemistry in 2014, where he majored in organic, inorganic, physical and analytical chemistry. He is currently completing a Ph.D. on the design and production of gold nanoparticles able to act as multimodal anticancer agents, being both drug delivery platforms and radiation dose enhancers.
Source: Read Full Article
